Background PCR has the potential to detect and precisely quantify specific DNA sequences, but it is not yet often used as a fully quantitative method. A number of data collection and processing strategies have been described for the implementation of quantitative PCR. However, they can be experimentally cumbersome, their relative performances have not been evaluated systematically, and they often remain poorly validated statistically and/or experimentally.
In this study, we evaluated the performance of known methods, and compared them with newly developed data processing strategies in terms of resolution, precision and robustness. Results Our results indicate that simple methods that do not rely on the estimation of the efficiency of the PCR amplification may provide reproducible and sensitive data, but that they do not quantify DNA with precision. Other evaluated methods based on sigmoidal or exponential curve fitting were generally of both poor resolution and precision. A statistical analysis of the parameters that influence efficiency indicated that it depends mostly on the selected amplicon and to a lesser extent on the particular biological sample analyzed. Thus, we devised various strategies based on individual or averaged efficiency values, which were used to assess the regulated expression of several genes in response to a growth factor. Quantitative PCR is used widely to detect and quantify specific DNA sequences in scientific fields that range from fundamental biology to biotechnology and forensic sciences.
For instance, microarray and other genomic approaches require fast and reliable validation of small differences in DNA amounts in biological samples with high throughput methods such as quantitative PCR. However, there is currently a gap between the analysis of the mathematical and statistical basis of quantitative PCR and its actual implementation by experimental laboratory users.
While qPCR has been the object of probabilistic mathematical modelling, these methods have not often been employed for the treatment of actual measurements. Therefore, the validity of the assumptions or simplifications on which these models are based is often unclear.
At the other extreme, the treatment of laboratory measurements is often fairly empirical in nature, and the validity or reproducibility of the assay remains usually poorly characterized from an experimental and/or theoretical basis. Thus, practical qPCR methods usually do not allow mathematically validated measurements, nor the determination of the statistical degree of confidence of the derived conclusions. Consequently qPCR results have been questioned , , with the consequence that semi-quantitative methods (e.g. End-point PCR) remain widely used.
Quantitative PCR amplifications performed in the presence of a DNA-binding fluorescent dye are typically represented in the form of a plot as shown in Figure, where the measured fluorescence is represented as a function of the PCR cycle number. An assumption that is common to all qPCR methods is that the fluorescence is directly correlated to the amount of double stranded DNA present in the amplification reaction. The amplification curves are sigmoid shaped and can be split into three phases. Reforestation tool eugene oregon.
Phase I (Figure ) represents the lag phase in which no amplification can be detected over the background fluorescence and statistical noise. This phase is used to evaluate the baseline fluorescent 'noise'. Phase II corresponds to the early cycles at which detectable fluorescence levels start to build up following an exponential behaviour described by the equation inserted in Figure. On a log scale graph, this corresponds to the linear phase, illustrating the exponential dynamic of the PCR amplification (Figure ).
During the later phase of the reaction, or phase III, the DNA concentration no longer increases exponentially and it finally reaches a plateau. This is classically attributed to the fact that one or more of the reactants become limiting or to the inhibition of amplification by the accumulation of the PCR product itself. Figure 1 Representations of real-time PCR amplification curves. The three phases of the amplification reaction are shown either on a linear scale (panel A) or on a semi-log scale (panel B). Panel A represents a typical amplification curve, while panel B depicts amplification curves generated from serial dilutions of the same sample, either undiluted or diluted 10- or 1000-fold (indicated as 1, 0.1 or 0.001, respectively). During the lag phase (phase I), the fluorescence resulting from DNA amplification is undetectable above noise fluorescence in part A, while in part B, some data points take negative values and are not represented.
This phase is used to evaluate the baseline 'noise' of the PCR amplification. Exponential amplification of the DNA is detected in phase II (cycles 16 to 23, panel A). This phase of the amplification corresponds to the linear portion of the curve in panel B (closed circles).
A threshold value is usually set by the user to cross the log linear portion of the curve, defining the threshold cycle value ( Ct). Phase II is followed by a linear or plateau phase as reactants become exhausted (phase III). The inserted equations describe the dynamic of the amplification during phase II. In a perfectly efficient PCR reaction, the amount or copy number of DNA molecules would double at each cycle but, due to a number of factors, this is rarely the case in experimental conditions. Therefore the PCR efficiency can range between 2, corresponding to the doubling of the DNA concentration at each cycle, to a value of 1, if no amplification occurs (Eq.
1 in methods). Furthermore, the efficiency of DNA amplification is not constant throughout the entire PCR reaction.
The efficiency value cannot be measured during phase I, but it may be suboptimal during the first cycles because of the low concentration of the DNA template and/or sampling errors linked to the stochastic process by which the amplification enzymes may replicate only part of the available DNA molecules. Quantitative PCR is used under the assumption that these stochastic processes are the same for all amplifications, which may be statistically correct for N 0 values that are large enough so that sampling errors become negligible. The efficiency reaches a more or less constant and maximal value that may approach 2 in the exponential amplification of phase II, and it finally drops to a value of 1 during phase III. This implies that any appropriate analytical method should focus on phase II of the amplification where the amplification kinetic is exponential. Therefore, the first step in any qPCR analysis is the identification of phase II, which is more conveniently performed when data are represented on a log scale (Figure ).
Another assumption of qPCR is that the quantity of PCR product in the exponential phase is proportional to the initial amount of target DNA. This is exploited by choosing arbitrarily a fluorescence threshold with the condition that it lies within the exponential phase of the reaction. When fluorescence crosses this value, the cycle is termed the 'Threshold cycle' ( Ct) or 'Crossing Point', and the higher the Ct, the smaller the initial amount of DNA. This is illustrated in Figure, which displays qPCR amplifications performed on serial dilutions of a cDNA sample. One of the first and simple methods to process qPCR data remains a set of calculations based solely on Ct values and is currently known as the Δ C tmethod ,. However, as such, this method assumes that all amplification efficiencies are equal to 2 or at least equal between all reactions.
Therefore it does not take into consideration possible variations of amplification efficiencies from one sequence or sample to the other. Thus, the Δ C tmethod may not accurately estimate relative DNA amounts from one condition or one sequence to the other. Consequently, other methods of data processing have been developed to estimate the efficiency of individual PCR amplifications –. Alternatively, amplification curves can be directly fitted with sigmoid or exponential functions (Methods section, Eq. 8) in order to derive the original amount of template DNA (Eq. Methods to estimate amplification efficiency can be grouped in two approaches, both of which rely on the log-linearization of the amplification plot.
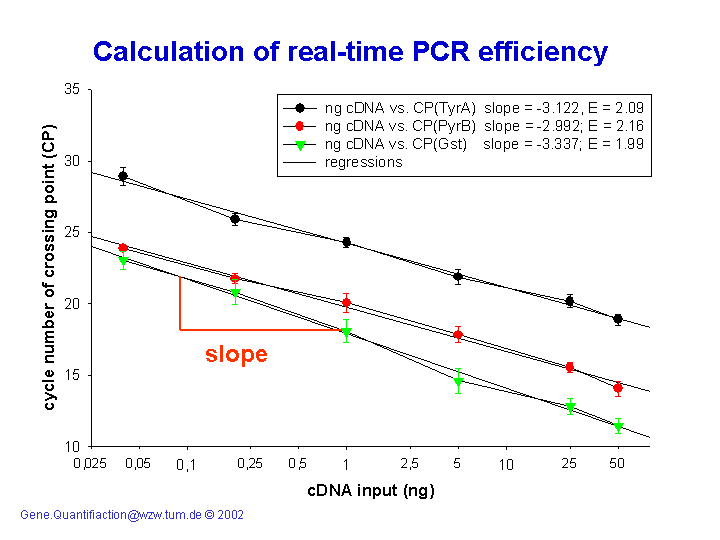
The most commonly used method requires generating serial dilutions of a given sample and performing multiple PCR reactions on each dilution ,. The Ct values are then plotted versus the log of the dilution (Figure ) and a linear regression is performed (Eq. 4) from which the mean efficiency can be derived (Eq. As stated above, this approach is only valid if the Ct values are measured from the exponential phase of the PCR reaction and if the efficiency is identical between amplifications. Figure 2 Measurement of the efficiency of a PCR reaction. A: Estimation of the efficiency using the Serial dilution (SerDil) method.
Five dilutions of a cDNA sample were amplified using the fibronectin (FN) amplicon. Each dilution was analyzed with five replicates PCR reactions and each data point represents one Ct value determined as in Figure 1B. Linear regression parameters and calculation of the efficiency value are shown in the inserted textbox. B: Screenshot of the LinReg PCR program analysis window, which allows the estimation of the efficiency value from each set of amplification curves 13. Data correspond to one of the reactions performed from the undiluted sample used in part A.
C: Comparison of the efficiency values obtained using the Serial dilution and LinReg methods for the FN amplicon. Efficiency values were determined from four independent cDNA samples using the Serial Dilution method as in part A, or the LinReg method as in part B. For each sample, efficiency value were either determined from one linear regression performed on 24 reactions altogether (Serdil) and error bars calculated from the standard deviation on the slope as determined from the linear regression method or the individual efficiency values determined from each of the same 24 PCR reactions (LinReg) were averaged, and error bars represent the standard deviation on the set of values.
The other method currently used to measure efficiency is based on Eq. 3, which associates an efficiency value with each PCR reaction.
This approach has been automated in different programs , one of which, termed LinReg PCR , was used in this study. LinReg identify the exponential phase of the reaction by plotting the fluorescence on a log scale (Figure ). Then a linear regression is performed, leading to the estimation of the efficiency of each PCR reaction. None of the current qPCR data treatment methods is in fact fully assumption-free, and their statistical reliability are often poorly characterized. In this study, we evaluated whether known mathematical treatment methods may estimate the amount of DNA in biological samples with precision and reliability. This led to the development of new mathematical data treatment methods, which were also evaluated.
Finally, experimental measurements were subjected to a statistical analysis, in order to determine the size of the data set required to achieve significant conclusions. Overall, our results indicate that current qPCR data analysis methods are often unreliable and/or unprecise. This analysis identifies novels strategies that provide DNA quantification estimates of high precision, robustness and reliability. Quantitative PCR usually relies on the comparison of distinct samples, for instance the comparison of a biological sample with a standard curve of known initial concentration, when absolute quantification is required , or the comparison of the expression of a gene to an internal standard when relative expression is needed. The equation inserted in Figure is used to calculate the ratio of initial target DNA of both samples (Eq. The error on the normalized ratio depends on the error on the Ct and the error on the efficiency, and it can be estimated from Eq.
However, the range and relative importance of the various components, and the origin of the error on practical measurements remain poorly characterized. To evaluate the reproducibility of Ct measurements and their associated error, we generated a set of 144 PCR reaction conditions corresponding to various target DNA, cDNA samples and dilutions (see Additional file for a description of targeted genes and amplicons). Each of these 144 reaction conditions was replicated by performing 4 or 5 independent PCR amplifications. This yielded a complete dataset of 704 amplification reactions which collection of raw data is given in additional file. Individual Ct values corresponding to each reaction conditions were averaged, providing a set of 144 Ct values and their associated errors. The standard deviation (SD) shows an increase of the error with higher Ct values, with SD values smaller than 0.2 for Ct up to 30 cycles, and spreading over 0.8 for Ct higher than 30 (Additional File ). Thus, all replicates with SD above 0.4 were excluded, which corresponds to some of the reactions with Ct above 30 in this study.
We conclude that Ct between 15 and 30 can be reproducibly measured leading to a dynamic range of 10 5, which is within the 4 to 8 logs dynamic range reported in other studies. In these conditions, Ct value determination is unlikely to be a major source of error when calculating normalized ratio of expression.
Thus, we then focused on the estimation of efficiency. Estimation of the efficiency of a PCR reaction We compared estimates of the efficiency obtained from two distinct methods: the generally used serial dilution (Figure ) and the alternative LinReg method (Figure ). With our experimental setup, estimation of the efficiency with the serial dilution method requires a set of 24 PCR reactions for a given sample and a given amplicon, using serially diluted template DNA. The efficiency obtained was compared to the average efficiency estimated from each of the reactions with the LinReg method. Efficiency estimates are comparable when looking at values given in Figure and, but they differed when comparing the efficiencies obtained from one of the four DNA samples (Figure ). Thus, we questioned whether the two methods provide statistically similar measures of efficiency, and whether they display similar reproducibility. The statistical equivalence of the LinReg and serial dilution methods was assessed using an analysis of variance (ANOVA, Table ), which indicated that the efficiency averages are not significantly different between the two methods except for one amplicon, corresponding to the Connective Tissue Growth Factor (CTGF) cDNA ( p.
Single one-way ANOVA was applied to the complete set of data to assess whether both the Serial dilution and the LinReg methods give similar averaged efficiencies ( H 0: efficiencies are equal, H 1: efficiencies are different): a p-value below 0.05 indicates that measured efficiencies are significantly different.F-test assessing whether the variance (error) induced by each method is the same H 0: variances are equal, H 1: var(serial dilution) var(LinReg): p. Next, we wished to determine which of the experimental variables may affect the precision of the estimation of efficiency. This was evaluated on the complete set of quantitative PCR reactions. Figure shows the distribution of the efficiencies measured for all reactions. Efficiencies ranged from 1.4 to 2.15 with a peak value around 1.85.
Theoretically, efficiencies can only take values between 1 and 2, and therefore they are expected to deviate from a normal distribution, as indicated by a Kolmogorov-Smirnoff test (not shown). However, the distribution appears to be sufficiently symmetrical to be considered normal, such that classical statistical tests can be validly performed. First, we determined if single PCR parameters (amplicons, cDNA samples, Ct value, etc.) may influence the efficiency value by performing a multiple ANOVA test on all values. The first four entries of Table indicate that the efficiency is most dependent upon the amplicon and relatively less on cDNA samples, as indicated by high F values, both of these effects being highly significant ( p.
A multiple ANOVA test was performed on efficiencies obtained from 700 reactions. Dependence with amplicon sequence, sample, Ct and dilutions were tested as well as all combinations of co-dependence. Df indicates the degree of freedom.
The larger the F-value, or the smaller the p value, the more significant is the effect of the corresponding parameter. Overall, these results indicate that efficiencies are highly variable among PCR reactions and that the main factor that defines the efficiency of a reaction is the amplicon. This is consistent with the empirical knowledge that primer sequences must be carefully designed in quantitative PCR to avoid non-productive hybridization events that decrease efficiency, such as primer-dimers or non-specific hybridizations. Efficiency might also depend upon the dilution for a minority of the cDNA samples, indicating that dilute samples should be preferred to obtain reliable efficiency values. DNA quantification models The models we evaluated in this study can fall into two different groups: being derived from either linear or from non-linear fitting methods. Comparison of qPCR data using models based on non-linear fitting methods (Eq. 8) is done simply by calculating the ratio of the initial amount of target DNA of each amplicon (Eq.
Calculating Pcr Product Size
9) as in the first part of Eq. The standard deviation of the ratio on a pool of replicate is calculated using Eq. Note that in this case, errors resulting from the non-linear fitting itself are not considered in the analysis. Linear fitting methods also allow the estimation of the initial level of fluorescence induced by the target DNA.
For instance, Eq. 3, upon which the LinReg method relies to determine efficiency, can also be used to determine F 0 as the intercept to the origin of a linear regression of the log of fluorescence. This figure can then be used to calculate relative DNA levels (Eq.
This calculation method was termed LRN 0. However, even small errors on the determination of the efficiency will lead to a great dispersion of N 0 values due to the exponential nature of PCR (Eq. Therefore, we considered alternative calculation strategies, whereby the efficiency is averaged over several reactions rather than using individual values, which should provide more robust and statistically more coherent estimations. We therefore evaluated the use of efficiency values calculated in three different manners. As the amplicon sequence is the main contributor to the efficiency, we used the efficiency averaged over all cDNA samples, dilutions and replicates of a given amplicon, as a more accurate estimator of the real efficiency than individual values. The error on the efficiency is no longer considered in the calculations of relative DNA concentrations, thus assuming that the estimator is sufficiently precise so that errors become negligible. This model is termed below ( PavrgE) Ct.
Alternatively, the small influence of the sample upon the efficiency was taken into account by averaging the efficiencies obtained for each dilutions and replicates of a given cDNA sample and a given amplicon. Thus, for a given cDNA sample and amplicon, one efficiency value is obtained from 24 PCR reactions. This value is used in further calculations, assuming again the average value to be a sufficiently good estimator of the efficiency so that the relative error may not be taken into account. This model was named ( SavrgE) Ct.Each line indicates how single efficiency values are grouped in each of the calculation models. For instance, the ( PavrgE) Ctmodel line indicates that one individual amplicon is chosen out of the 7 available ones, then efficiencies estimated from the different cDNA samples, dilutions and replicates are all pooled, leading to the determination of one efficiency value from 100 reactions. In the ( SavrgE) Ctmodel, individual efficiency values are calculated for each cDNA sample and each amplicon averaging efficiency values from the 5 replicate reactions performed on the 5 dilutions (25 values overall). Evaluation of the quantitative PCR calculation models.
Calculating Pcr
The dilutions of a given sample form a coherent set of data, with known concentration relationships between each dilution. Each calculation model was therefore used on each dilution series, using the undiluted sample for normalization.
All data can be presented as measured relative concentrations, the undiluted dilution taking the relative concentration value of 1, the 10-fold dilution taking the value of 0.1, the 50-fold dilution a value of 0.02, and so on. The measured relative concentrations for all dilutions, samples and primers and the associated errors were calculated using each model from the complete dataset of 704 reactions. For the models giving a direct insight to the initial N 0 values, N 0 were averaged for each amplicon and cDNA sample, and they were plotted in comparison with the expected concentrations relative to the undiluted samples (Figure ). The models were evaluated on three criteria: resolution, precision and robustness. Figure 4 Comparison of the different calculation models when applied to samples of known relative concentrations. Each cDNA samples serial dilutions were processed with the indicated models, measured concentration were expressed as relative to the undiluted sample. Then results of all amplicons were averaged for a given model.
We defined the resolution as the ability of a model to discriminate between two dilutions. Relative concentrations were compared pair-wise between adjacent dilutions. Typically, it can be seen in Figure that models did not give uniformly coherent results. For instance, models that do not rely on explicit efficiency values, such as the sigmoid or exponential models, are unable to discriminate between the 0.1 and 0.02 relative concentrations, which shows a lack of resolution in this range of dilutions. The Δ Ct, ( PavrgE) Ctand ( SavrgE) Ctmodels performed well under this criterion, allowing easy discrimination of the 10-fold and 50-fold dilutions in this example. The resolution was statistically evaluated with a coupled ANOVA-LSD t-test, which is a two step analysis of variance (ANOVA) coupled to a t-test run under the Least Significant Difference method (LSD). Unsurprisingly, the ANOVA test indicated that for all models at least one of the measured concentrations differed significantly from the others as expected (data not shown).
To further assess if all measured concentrations significantly differ from one another, or if some are undistinguishable, a coupled t-test was performed on pairs of adjacent dilutions in a given serial dilution series. Results are summarized in Table. All models were able to discriminate the undiluted condition from the 10-fold dilution (highly significant, p. Relative concentrations Δ CT E Ct (Savrg E) Ct (Pavrg E) Ct LR N 0 Sigmoid Exponential t-value p-value t-value p-value t-value p-value t-value p-value t-value p-value t-value p-value t-value p-value 1 – 0.1 122.7 0.000 18.6 0.000 141.7 0.000 141.3 0.000 18.9 0.000 29.1 0.000 18.2 0.000 0.1 – 0.02 7.3 0.000 1.9 0.030 12.6 0.000 12.6 0.000 3.5 0.000 -4.4 N/A 0.8 0.225 0.02 – 0.01 0.7 0.232 0.4 0.360 1.6 0.052 1.6 0.052 0.9 0.190 6.1 0.000 -0.4 N/A 0.01 – 0.001 0.5 0.325 0.3 0.375 1.3 0.105 1.2 0.107 0.5 0.306 0.1 0.478 2.7 0.004. A ANOVA coupled t-test was performed to determine which dilutions are statistically different from one another.
The comparison of adjacent dilutions are shown. A p-value higher than 0.05 indicate that the model is unable to discriminate between the two adjacent dilutions. N/A stands for not available and indicate that the expected higher concentration of the comparison was in fact calculated to be lower from the qPCR results. The precision of a model is defined by its ability to provide expected relative concentrations of the known dilutions. Again Figure shows that the ( PavrgE) Ctand ( SavrgE) Ctmodels provide precise relative concentration values over all dilutions, with the measured relative concentrations matching the expected ones. Estimations obtained by the Δ Ct model appear to be less reliable, with a systematic under-representation of concentrations. This result is expected since all of our amplicons have efficiencies that are below 2 (see Additional File ).
We statistically evaluated the precision of each model by plotting the expected relative concentration against the measured relative concentration averaged from all primers and samples (Additional File ). A linear regression was done on the data obtained from each model and a t-test was performed to determine if the slope is statistically different from 1. A low p-value in Table is associated to a high probability that the slope is different from 1, indicative of a poor correlation between expected and measured values. As before, the ( PavrgE) Ctand ( SavrgE) Ctmodels outperformed all other models, being more precise than the and sigmoid models, the exponential, Δ Ct and LR N 0 displaying lowest precision. Expected and measured relative concentrations were plotted and a linear regression was performed (Additional File ). The slopes values were submitted to a t-test to evaluate if they are statistically different from 1, which would indicate a poor correlation between expected and measured concentrations. A p-values smaller than 0.05 indicate that the slope is significantly different from 1.
Correlation coefficient (r 2) values indicate the width of the spreading of individual data around the linear regression line. Higher r 2 indicate more robust models. Finally, the robustness is related to the variability of the results obtained from a given model, and it indicates whether trustable results may be obtained from a small collection of data. For instance, a model could be very precise (eg providing a slope of 1) with a large data set, but the distribution of the points around the regression line could be very dispersed. Such a model would not be robust as a small data set would not allow precise measurements.
Thus, the robustness of a model was estimated from the standard deviation of the slope and the related correlation coefficient of the linear regression ( r 2), with higher r 2 values indicating more robust models. Three models showed high robustness, the Δ Ct, ( PavrgE) Ctand ( SavrgE) Ct, followed by E Ct(Table ). Overall, only two calculation models combine high resolution, precision and robustness, namely the ( PavrgE) Ctand the ( SavrgE) Ctmethods. However, only the slope of the ( SavrgE) Ctdid not statistically differ from 1. Model evaluation on a biological assay of gene expression r.
Are optimized for maximum performance and minimal environmental impact. Kits are available for plasmid miniprep, gel extraction, PCR & reaction cleanup, and total RNA purification. For maximum convenience and value, columns and buffers are also available separately. You’ll be Thrilled to Pieces Do you need a faster, more reliable solution for DNA fragmentation and library construction? Our new with novel fragmentation reagent meets the dual challenge of generating high quality next gen sequencing libraries from ever-decreasing input amounts AND simple scalability.
Are optimized for maximum performance and minimal environmental impact. Kits are available for plasmid miniprep, gel extraction, PCR & reaction cleanup, and total RNA purification. For maximum convenience and value, columns and buffers are also available separately. You’ll be Thrilled to Pieces Do you need a faster, more reliable solution for DNA fragmentation and library construction? Our new with novel fragmentation reagent meets the dual challenge of generating high quality next gen sequencing libraries from ever-decreasing input amounts AND simple scalability. PCR Protocol for Taq DNA Polymerase with Standard Taq Buffer (M0273) Protocols.io also provides an where you can discover and share optimizations with the research community. Overview PCR The Polymerase Chain Reaction (PCR) is a powerful and sensitive technique for DNA amplification (1).
Taq DNA Polymerase is an enzyme widely used in PCR (2). The following guidelines are provided to ensure successful PCR using NEB's Taq DNA Polymerase. These guidelines cover routine PCR. Amplification of templates with high GC content, high secondary structure, low template concentrations, or amplicons greater than 5 kb may require further optimization. Protocol Reaction setup: We recommend assembling all reaction components on ice and quickly transferring the reactions to a thermocycler preheated to the denaturation temperature (95°C). Component 25 μl reaction 50 μl reaction Final Concentration 10X Standard Taq Reaction Buffer 2.5 μl 5 μl 1X 10 mM dNTPs 0.5 µl 1 μl 200 µM 10 µM Forward Primer 0.5 µl 1 μl 0.2 µM (0.05–1 µM) 10 µM Reverse Primer 0.5 µl 1 μl 0.2 µM (0.05–1 µM) Template DNA variable variable.
When I left off my last discussin of tboot PCR calculations I gave a quick intro but little more. In this post I’ll go into details for calculating the first of them: PCR17. There have been a number of discussions with regard to calculating or verifying PCR values on the tboot-devel mailing list and they were extremely useful in writing this code and post. These all fell a bit short of what I wanted to accomplish in that all approaches extracted hashes from the output of the txt-stat program (the tboot log) and used those hashes to re-construct the PCR values. I wanted to construct all hashes manually, to measure and account for the actual things that TXT and tboot were measuring and storing into the PCRs and to do this independent of an actual measured launch. Basically this translates to isolating the things being measured, extracting them (if possible) and use them to reconstruct the PCR value on any system, like a build server or an external verifying party.
The process is pretty straight forward, though time consuming, and the specification is phrased in such a way as to force some guess and check. There’s even a bit of a trick in the end which requires that we go digging around in the tboot source code which is always fun. I’ll also present a bit of code that will automate the calculation for you so if you’re anxious and don’t want to read any more you can go straight to the code which can be found here:. DISCLAIMER: The code in the pcr-cal git repo is very much a work-in-progress and should be considered unstable at best so YMMV. The spec that defines the DRTM specific PCRs is the “PC Client Implementation for BIOS”. These are PCRs 17 through 20.
Calculating Pcr Products
Their individual use however is hardware specific and on Intel hardware, the definitive source of data on what gets extended into which of these PCRs as part of establishing a DRTM is a document titled “Intel® Trusted Execution Technology (Intel® TXT) Software Development Guide: Measured Launched Environment Developer’s Guide”. Quite a mouth full. Anyways section 1.9.1 covers PCR17 but the details of what various bits are measured are spread out over the document. A default tboot configuration will cause 3 extends to this PCR so we’ll break this post up into 3 sections, each one describing the hashes that go into the 3 consecutive extend operations. First extend: SINIT ACM The first thing that’s extended into PCR17 is the hash of the SINIT ACM. This is a binary blob that Intel ships which is used by tboot to establish the DRTM on a platform.

The binary code in the ACM is chipset specific so there are a number of ACMs out there to chose from. Tboot automates the process so if you’re unsure which ACM is the right one for your platform you can configure your bootloader to load every ACM and tboot will pick the right one.
This will slow your boot process down considerably though and selecting the proper one isn’t hard with a bit of reading so don’t be lazy. With the right ACM in hand you’d think it would be a simple matter of calculating the sha1 hash of the file and extending that into PCR17. That’s not the case though. There are two little details that must be sorted first.
Case Of merupakan salah satu pernyataan kondisi dimana pernyataan lebih sering digunakan untuk menjalankan beberapa program yang telah digabung/ dipadukan menjadi satu program. 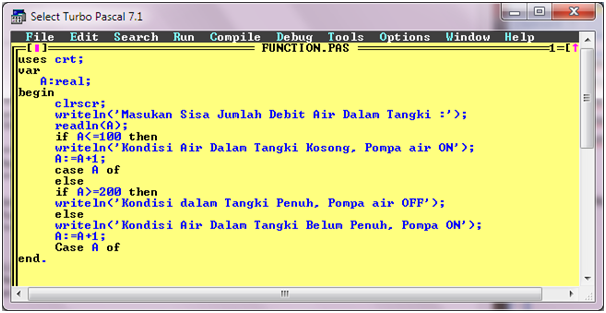

Depending on the version of the ACM you’re using the hash algorithm may be sha1 or it may be sha256. ACMs version 7 or later will use sha256, while earlier versions will use sha1. The current version of the ACM format is 8 so most modern hardware will need a sha256 hash (not to mention that most OEM implementations of TXT 3 years or older never worked in the first place snap!). Further, there are some fileds in the ACM that aren’t included in the hash. The logic behind this escapes me but the apendix A.1.2 specifies that some fields are omitted. Quoting the spec: “Those parts of the module header not included are: the RSA Signature, the public key, and the scratch field.” That sounds like 3 fields from the ACM right?
Wrong: there’s a 4th field omitted as well and that’s the RSA exponent. I guess they meant for the exponent to be included in the definition of “public key”? Thanks for being explicit. Anyways omit the fields: RSAPubKey, RSAPubExp, RSASig and the Scratch space, got it.
To omit these fields from the hash we’ve gotta parse the ACM. I’ll cover this code at the end. Finally the 32 bits that make up the EDX register which hold the flags passed to the GETSECSENTER instruction are appended to the hash of the ACM. We represent PCR17 at the first extend operation thusly.
PCR171 = Extend(PCR170 SHA256 (ACM EDX)) PCR171 = Extend(PCR170 SHA256 (ACM EDX)) where PCR170 is the state of PCR17 at time = 0. PCRs are initialized to 20bytes of 0’s so PCR170 is 20 bytes of 0’s. Second Extend: Heap Data The second extend to PCR17 includes various bits of data from the TXT heap. Appendix C describes the TXT heap as a contiguous region of memory set asside by the BIOS for use by ‘system software’ (aka BIOS) to pass data to the SINIT ACM and the MLE.
PCR17 is extended with the sha1 hash of either 6 or 7 concatenated pieces of data depending on the version of the ACM. The following fields are concatenated together and their sha1 hash is extended into PCR17 for the second extend:.
BiosAcmId. MsegValid. StmHash. PolicyControl. LcpPolicyHash. OsSinitCaps or 4 bytes of 0’s as specified by the LCP (more on this next) If the SINIT to MLE data table version is 8 or greater an additional 4 bytes are appended representing the processor S-CRTM status.
These 4 bytes are in the ProcScrtmStatus field in the SINIT to MLE data table. The second extend to PCR17 could be represented as follows for SINIT to MLE data table versions. $ txt-heapdump -pretty -i txtheap.bin $ txt-heapdump -pretty -i txtheap.bin Once you have the heap as a file you can use the pcr-calc library to parse and extract various bits.
Again, I’ll present the utility that does it all for you at the end. But first, the third and final extend Third Extend: Launch Control Policy (LCP) You’d expect that all of the values that are hashed as part of PCR17 are discussed in the spec under section “PCR 17” and you’d be wrong. A couple of sections deeper where the LCP is discussed, you’ll find a description of how the LCP policy is measured and it turns out that this measurement gets extended into PCR17 as well! There are a number of rules laid out in this section for how the system behaves when there’s no ‘Supplier’ or ‘Owner’ LCP present. Specifically the spec states: As a matter of integrity, the LCPPOLICY::PolicyControl field will always be extended into PCR 17. If an Owner policy exists, its PolicyControl field will be extended; otherwise the Supplier policy’s will be. If there are no policies, 32 bits of 0s will be extended.
I’ve not gone through this section with a very thorough eye so I’m not an authority here, but tboot seems to ignore these rules and instead loads a default policy when there isn’t one in the TPM NV RAM. Not saying this is good or bad, right or wrong, just pointing out that this is what tboot does and it was something that I had to figure out in order to calculate the value of PCR17 independently on my test systems. So my goal here is to calculate PCR values. If your system is like mine and both you (the ‘Owner’) and the ‘Supplier’ (your OEM) didn’t provide an LCP, how do we measure the default policy from tboot? The only thing I could come up with is to pull apart the tboot code and copy the hard-coded structures into a C program and then dump them to disk in binary.
The hash of this file is the one we need to extend into PCR17 along with the LCP PolicyControl value. I’ve added a class to the pcr-calc library to parse the necessary parts of the binary LCP to support this operation. The program that dumps the binary LCP from tboot is: lcpdef I’ve kept this utility in the pcr-calc project to reproduce the LCP on demand. I considered only keeping around the LCP binary in a data file but in the event that the default tboot policy changes in the future I wanted to keep the program around to dump the binary structures. When executed this program just dumps the binary policy so you’ll have to redirect the output.
Bio572: The Polymerase Chain Reaction Polymerase Chain Reaction Amplification In the polymerase chain reaction, a DNA template is repetitively:. denatured into single stranded molecules,. annealed to specific oligonucleotide primers (one specific primer per strand),. copied with DNA polymerase to extend the primers to the end of the DNA strand.
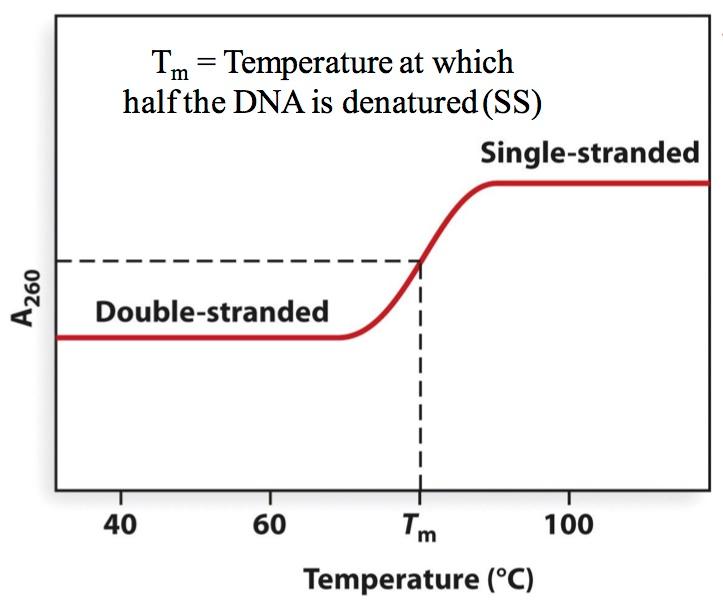
The reaction-in brief Step 1: Denaturation (95 oC-98 oC) Why must the DNA be denatured into single strands? Because without separation of strands, you would not be able to anneal (i.e. Hybridize) specific primers in the next step. Step 2: Annealing (45 oC-65 oC) Primers in excess The annealing reaction is very efficient because the primers are 'in excess' in the reaction. In a typical PCR reaction, 10,000 molecules of a template may be used, which is 1.6 x 10 -20 moles (0.016 attomoles). On the other hand, 5 picomoles of each primer may be used (5 x 10 -12 moles) - that is a 3 x 10 8 fold excess. Temperature controls annealing rate The rate of annealing is controlled by adjusting the temperature of the solution.
At 55 C under most PCR salt conditions, typical primers of 18 nt. In length efficiently hydrogen bond to a DNA template. Adjustments in the protocol are made to account for the G/C vs. A/T richness of the primer and the overall length. There are many programs or Web sites at which one may calculate the Tm based on sequence and salt condition.
Step 3: Extension (65 oC-75 oC) You can see, from a comparison of the figures for step 1 and step 3, that we now have two double-stranded DNA copies of the sequences between the specific primers. By denaturing these two copies and repeating the annealing and synthesis steps, we can obtain four copies. Repeat Now if we repeat the process again, we can obtain eight copies.
Note that the sequence between the two primers is being copied or 'amplified' exponentially, whereas the original template is not. Many of these copies have 3' overhanging ends, because the primer sequences are only extended in one direction (synthesis is only 5' to 3'). These longer versions are generated only from the original template, and not from the copies; as a result they are not generated at an exponential rate. On the other hand, the shorter versions (as in the bottom two molecules in the figure above) contain copies of the DNA between the two primer sequences, and are 'amplified' at an exponential rate (1, 2, 4, 8, 16, 32, 64, 128, 256, 512, 1024, and so forth). If things worked perfectly, we could obtain approximately a 1000-fold amplification for every ten cycles of synthesis!
An overview of what is needed: A pair of short oligonucleotide primers specific for a DNA sequence, with the ability to hybridize to the opposite strands of that molecule (3' ends pointing 'towards' each other):. A DNA template.
A thermostable DNA polymerase (such as Taq or Pfu polymerase), and all four dNTP substrates (meaning dATP, dGTP, dCTP, dTTP). A machine that can change the incubation temperature of the reaction tube automatically, cycling between approximately 98 C (for denaturation), 55 C (for oligonucleotide annealing), and 72 C (for synthesis). The temperature changes: When you program the thermocycler, you specify a series of temperatures and times, such as: Temperature Time 98 C 30 seconds 55 C 30 seconds 72 C 60 seconds.and specify the number of times this series should be repeated (for example, 35 times). Multiple program segments may be linked together, as in the following example: Temperature Time program segment 1, do 1 time: 98 C 5 minutes program segment 2, do 35 times: 98 C 30 seconds 55 C 30 seconds 72 C 60 seconds program segment 3, do 1 time: 72 C 10 minutes program segment 4, do 1 time: 4 C 999 minutes What was the purpose of each of these segments? Program segment #1: To denature the template fully, prior to the first synthetic step. If this step is excessive, there can be damage to the enzyme or template, reducing the efficiency.
If too short, the template is not available for synthesis. Program segment #2: To amplify the DNA fragment, with the following steps taken:. Denature DNA at 96 C. Anneal oligonucleotides at 55 C. Synthesize DNA at 72 C The individual temperatures (96, 55, 72) may be optimized for each reaction, however the thermostable polymerases generally work well at 72-74 C, and typical oligonucleotides anneal well at 45-65 C. Program segment #3: Finish synthesis of any partially completed fragments.
Program segment #4: Cool samples while waiting for researcher to finish nap. In this type of program, the temperature changes are as rapid as the machine can manage, usually taking 30 seconds to a minute to complete.
Some advanced machines can change the temperature between these steps in just seconds, and these speed up the PCR process considerably. This type of temperature profile could be represented by a square wave plot. Step cycle file There are times when you don't want the temperature changes to be rapid, and here's an example: Suppose we are trying to work out the conditions for a polymerase chain reaction using two degenerate oligonucleotides that are approximately 1000-fold degenerate. We would like to use that handy web site to determine the Tm, so we would know what annealing temperature to program into the machine, but we don't actually know which of the 1000 versions of each oligonucleotide will be an exact match to the target sequence. What we would really like to do is to introduce some flexibility into the temperature cycle, so that every potential oligonucleotide has a fair chance of annealing to the target. What do we do? Answer: We program the temperature cycler so that it gradually changes the annealing temperature, thereby exposing the reaction to a range of temperatures.
In this kind of program (characteristic of File #3 in the Perkin Elmer 480 instrument), the timing at each temperature and between each temperature are specified. Rapid changes can be programmed by setting a 'between temperature' time of only one second (it obviously takes longer than that to change temperatures, so the machine just does its best). In this example, the temperature gradually increases from 55 C to 65 C over a 1 minute period. Thermocycle file Otherwise, you will notice how similar it is to the square-wave version described before. If you've got your MasterCard ready, here are some of the models (past and present) from the Perkin Elmer showroom: The machines have a metal 'hot block' that conducts heat to the samples, and a heating/cooling fluid that circulates through the inside of the metal block to change the temperature. A cover protects you from burning yourself on the block.
Actual yield is less than the theoretical maximum PCR is usually represented by the maximal theoretical yield, which is to double the amount of product every cycle. In practice you do not achieve that level of synthesis. Here is an example of synthesis specifications from: Using the DNA Thermal Cycler 480 and the GeneAmp® PCR, an amplification yield of at least 100,000 fold of the Lambda Control DNA target can be achieved with: 0.2 µM each of the Lambda Control Primers. 0.1 ng of Lambda Control DNA target. 100 µL reaction volume with a 50 µL mineral oil overlay in a 0.5 mL GeneAmp® Thin-Walled Reaction Tube.
25 thermal cycles: 94 degrees C for 1 minute and 68 degrees C for 2 minutes. An amplification yield of 100,000x after 25 cycles would mean at each cycle 1 template would yield 1.58 templates for the next round of synthesis. How was this calculated? If c is the number of copies made per round of synthesis then c 25 = 100,000 = 10 5 so c 5 = 10 and so 5(log c) = log 10 = 1 so log c = 0.2 and c = 1.58 (approximately) (Or you could calculate the 25th root of 100,000 on a calculator, if you prefer.) If we obtain 1.58 copies instead of the theoretical maximum of 2 copies, then the efficiency of the reaction could be said to be 79% (because 1.58/2.00 = 0.79).
One reason this calculation is important, is that a slight loss of efficiency is magnified through the amplification. A reaction may appear to have not worked if the efficiency drops (in each cycle) by just a few percent. Optimization is critically important in the polymerase chain reaction. Specificity problem The appropriate annealing temperature can be calculated from the base sequence and length, noting that longer oligonucleotides can form more hydrogen bonds with a target and therefore have a higher annealing temperature. Similarly, the fraction of G or C nucleotides in an oligonucleotide affects the annealing temperature because GC base pairs form three H bonds and AT base pairs form only two.
If there is any degeneracy or mis-match between the oligonucleotide and the target, the annealing temperature will be lower. A typical annealing temperature one might use is 50 to 60 degrees Celsius. There is a problem with using annealing temperatures lower than that, because nonspecific products may be amplified.
These are side reactions in which an oligonucleotide may form just a few Watson Crick base pairs with a template, and are usually unwanted. Here is an example: GGATAGGACCTAGGAGGACCAGGAGATCCCGCCTACCGAAGGACG-3' synthesis.
